Genetic Variability by DesignJournal of Creation 18(2) 2004 Creation Ministries International Download PDF File (printable version)
Watch Seminar Video Abstract Although most genes remain unchanged from one generation to the next, others are highly variable (hypervariable) in comparison, and new alleles are accumulating rapidly in living populations. Cellular mechanisms have not been adequately sought to explain the intentional production of these changes, but it is becoming clear that homologous recombination is involved. Since its discovery during meiosis, these reactions were assumed to occur randomly along the length of chromosomes, and only involved with gene crossovers. It is now well known that meiotic recombination is not the random process it was originally assumed to be, and controlled by highly organized regulatory systems. In addition, a form of homologous recombination has been discovered that is responsible for creating diversity in variable genes, and was recently linked to single base-pair substitutions in immunoglobulins. New allele formation may indeed be the key to explaining the rapid production of distinct breeds, but their presence in the genome has been assumed the result of random mutations. Therefore, the ability of the cell to purposefully edit genes requires evaluation. Introduction There are two conceptual models that overshadow current theories related to the production of diversification within organisms. Darwinian evolutionists propose that genetic variability is the result of random mutations that have accumulated gradually over millions of years. In contrast, creationists generally assume that God made all useful genetic information during the creation of the world. Both largely agree that groups such as the Family Canidae evolved from a common ancestor, and the rapid development of variety during the domestic dog breeding, was simply the result of shuffling preexisting genes. This common genetic theory fails to explain the rapid production of the significant diversity when presented with the assertion that the wolf was a pure breed while in nature. A purebred organism is a genetic homozygote for the characteristics in question, and will therefore pass its traits to every offspring produced. In other words, wolves only give birth to wolf pups, but from them a tremendous genetic and morphologically diverse number of dogs have been isolated in a relatively short period of time. Explaining this conundrum cuts to the very heart of the creation vs. evolution controversy. The canine baramin (Biblical kind) likely speciated rapidly following the flood of Noah during geographic radiation, and each new population was then naturally selected until varieties such as the wolf, fox, hyena, jackal, etc., each became purebred. Selection is the process of inbreeding specific genes from a diverse genotype until alternate variations of genes (alleles) are eliminated, and the bloodline will only then breed true. As a result, the purebred organism is genetically limited, and in many respects the wolf should be considered no different in its ability to produce offspring diversity than any of the dog breeds we have today. Although speculative for some time, genetic studies have now thoroughly established that dogs were bred from domesticated gray wolves.1 It is also a matter of historic fact that most breeds have been selected to purity within the last few hundred years. However, when compared to the wolf, the domestic dogs are found to have divergent sequences not possessed by their progenitors. Due to these findings some geneticists now believe that wolf domestication must have begun more than 100,000 years ago despite the fact that archaeological findings can not verify their existence beyond 14,000 years.2 If one believes that the Biblical flood of Noah was a recent and global event, then another problem presents itself. Genes have fixed locations within the genome called loci, and sexually reproducing organism only carry two alleles (variation of a gene) per locus.(Figure 1) If the number of breeding pairs released from the ark can be taken as accurate data for this recent genetic bottleneck, then the maximum number of original alleles is known. According to the Bible, no more than seven breeding pairs were preserved for each baramin, and these 14 individuals can only carry 28 alleles per locus. Since only 8 people were alive following the flood, there should be a maximum of 16 alleles at any given locus in humans. However, it is now clear that many genes exist within populations as hundreds or even thousands of alleles. For example, 240 alleles have already been discovered in the human HLA-B locus. In spite of such concrete evidence, most creationists still tend to assume there is no mechanism for generating new genetic information,3 and indeed the following quote may be exemplary of the current state of thinking regarding intelligently designed genetic heredity. "Recombination explains why children look different from their parents. This shuffling of the genes can produce superior combinations of different genes. However, because we see that mutations are incapable of supplying useful variation, the useful genes that are there to be shuffled must have been created at the beginning."4 The History of Life. Lane P. Lester. Creation Research Society Quarterly 31(2) 1994 p96.
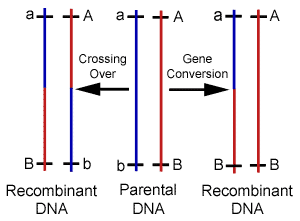
Mutation vs. Recombination There are two potential sources for changes to gene sequence; mutations and recombination. The cell recombines DNA for various reasons including the purposeful generation of diversity. Mutations on the other hand are changes resulting from of exposures to foreign mutagens, or the result of errors during biochemical reactions such as DNA replication. Changes to genes are almost universally attributed to the latter, however, replication attempts to copy the genome verbatim, while recombination is intentionally making alterations in a largely uncharacterized manner. Therefore, any changes found should be automatically assumed the result of recombination. This simple logic has escaped modern philosophers who do not recognize the existence of cellular design behind such variability. Although it is certainly possible for mutations to produce a beneficial change to the genome, finely tuned environmental adaptations are not likely accomplished by randomly altering genetic code. Because the production of variation during domestic breeds occurs over an extremely short period of time, it is obvious that recombination produces these changes. Nevertheless, mutations that have accumulated over millions of years are believed by evolutionists to have created the alleles responsible for the physical differences between breeds. Naturalism assumes that life and genetic information exist without intent, and therefore, the changes to genes that ultimately drive evolution are presumed to be independent of cellular intent. Darwinian evolution essentially requires mutations as the source of new genetic information to explain the existence of variability before cellular mechanisms developed. Due to this theoretic necessity, cellular reactions have not been adequately sought as a purposeful source of new alleles. It should be noted that the mitochondria genome previously served as the only evidence to support gene variability independent of recombination because the organelle was thought to originate exclusively from maternal contributions. However, it was recently reported in Science that recombination between parental DNA also occurs within the mitochondria. Mixing of paternal with maternal mitochondria sequences was recently found, and the investigators have concluded that recombination occurred between the organelles following fertilization.5 The mechanisms responsible for the large pools of alleles found within populations today are almost entirely theoretic. Following cell division, we simply can not determine whether mutations or recombination were responsible for any particular genetic alteration. But, since many alleles must have accumulated rapidly if the young earth position is correct a mechanism should be sought that is able to introduce these alterations in a controlled and systematic fashion. To this point, recent discoveries have shown that many genes are highly variable or polymorphic in comparison to others, and possess regions that change significantly from one generation to the next. Although changes to gene sequence are still commonly assumed the result of mutations, evidence has surfaced which is helping to demonstrate that new alleles are the result of purposeful genetic recombination. Characterizing the Various Functions of Homologous Recombination Homologous recombination (HR) is the name given to a large group of reactions during which the cell uses a stretch of DNA to alter a similar (homologous) sequence for several purposes including repair and general editing. The cell's intention in performing these manipulations has been preprogrammed by the creator in much the same way as we program a computer to perform functions independent of further human input. Offspring are always genetically unique due to HR, but we have only been able to recognize the most obvious products and the desired outcomes remain theoretic.(Figure 2)
Our knowledge of HR comes predominantly from the bacteria E. coli, and its effect during sexual reproduction (meiosis) has been studied mostly using lower eukaryotes such as baker's yeast, as well as fruit flies. Recent work with mice has provided additional information from mammals, and shown that substantial differences exist between unicellular and multicellular organisms. However, as with most cellular housekeeping mechanisms, the basic details and many genes involved in HR appear conserved among the multitude of life forms on earth.6 It is now widely recognized that genetic editions through HR are part of a highly coordinated process involving a cascade of specific macromolecule interactions,7 and controlled by highly organized regulatory systems.8 In particular, the induction of recombination during meiosis is reliant upon several genes, and is regulated by a complex network of cell signaling mechanisms.9 Since their discovery and use in the construction of genetic maps, it was assumed that gene crossovers during meiosis occurred at random intervals along chromosomes. It was believed that the frequency of gene crossovers was directly related to the distance between genes, but a variety of discoveries have illustrated the existence of differential recombination rates and patterns, and forced a revision of map distances. It is now a well-known fact that recombination frequency is not constant in any one particular cell. Reactions occur more frequently in some regions of the genome than in others with variations of several orders of magnitude observed. These hyperactive regions have been termed as "hot spots" as opposed to inert "cold spots" where little to no exchange is found.10 The frequencies of recombination events are also nonrandom. The rates are found to be significantly higher when comparing germ-line with somatic cell types. For example, mitotic recombination frequencies in the fungus Ustilago maydis have been estimated at 2.9 x 10-7; whereas, in meiosis the rates are closer to 1.9 x 10-3. Sex-specific differences in recombination frequency have also been elucidated. Standard linkage analysis was used to confirm that females have a higher recombination rate than males, and males recombine preferentially in the distal regions of the chromosome. These and other techniques were also separately used to establish the existence of significant inter-individual variation in recombination over short intervals.11 Still other researchers have demonstrated background effects on the frequency of recombination using immunostaining techniques to assess meiotic exchange patterns. It has now been found in many cases, that crossover events are non-randomly distributed and display positive interference.12 In addition to exchanges during cell division, HR is involved with many other forms of genomic DNA editing. For example, recombination is induced or shut off as a preprogrammed cell function during differentiation and development. It is also used to perform error-free DNA repair, which in this case serves to prevent unintentional variability. In fact, HR maintains the integrity of the genome through the correction of several different types of DNA damage.13 Homologous recombination is stimulated by double-stranded breaks during any stage of the cell cycle, and is also responsible for performing deletions, duplications, and translocations between dispersed homologous, which are frequently a response to stress.14 The specific details or exact sequence homology required for HR remain largely unknown, but the plethora of functions accomplished by these reactions has elevated them to the position of master mechanic responsible for virtually all forms of sequence editing and maintenance. There is an interesting new class of HR only recently recognized that shares common mechanisms with meiotic crossovers, and is likely responsible for the formation of new alleles. The process known as gene conversion uses template DNA to edit active sequences. During this process, pseudo genes previously referred to as junk DNA is frequently used to make these changes.15 Gene conversion can be easily distinguished from crossovers in most cases because only one of the homologues are altered.(Figure 3) It has now been thoroughly documented that mitotic recombination via gene conversion is able to create genetically altered cells, and researchers have suggested that this process can generate a gene with novel functions by rearranging various parts of the parental reading frames.16 DNA is also repaired through conversion when an intact copy from the sister chromatid or homologous chromosome is used to replace the damaged region. Gene conversion is now understood to be responsible for performing many alterations that were previously attributed to mutations or other repair mechanisms.
Hypervariable Genes Diversification within a population occurs because the genes involved with the production of characteristics exist as a variety of alleles, and therefore traits are polymorphic or available in more than one form. Closely related species are commonly found with extremely high numbers of alleles. For example, the cystathionine ß-synthase gene locus has been intensely studied in humans, and Exon 8, in particular, has a high frequency of single nucleotide alterations. It is estimated that approximately 5% of human Caucasians possess variations in this region.17 Evolutionists generally assume that new alleles are the result of random mutations that have accumulated gradually over millions of years. However, living populations have been tested only decades following severe genetic bottlenecks to find surprisingly high genetic diversity. This strongly suggests a mechanism for rapidly restoring variability, and the yet this possibility has not yet been adequately explored.18 However, an explanation for this continued diversity was suggested when it was discovered that many genes in every genome are highly diverse (hypervariable) in comparison to others. Not all genes are variable. The majority of genes in the genome are involved with housekeeping functions, and are commonly found unchanged even when comparing vastly different organisms. In contrast, variable genes change significantly from one generation to the next and show nonrandom patterns within any given gene.19 The characterization of variable genes to date suggests overwhelmingly that this diversity is systematically produced through gene conversion while under tight cellular control. For example, variable genes have hot and cold spots of activity similar to those found among gene crossovers in meiosis.20 They also frequently have greater diversity than the neutral regions between reading frames.21 It has likewise become evident that variable genes retain codons at specific locations within the variable region.22 A preponderance of non-synonymous substitutions over synonymous has provided even further evidence against randomness.23 It is becoming increasingly questionable that variability is the result of random mutations as commonly claimed by evolutionists. The following quote adequately validates this assertion. "Adaptive evolution has long been regarded as the result of postmutational sorting by the process of natural selection. Mutations have been postulated to occur at random, producing genetically different individuals that then compete for resources, the result being selection of better adapted genotypes. Molecular biology has demonstrated, however, that the rate and spectrum of mutations is in large part under the control of genetic factors. Because genetic factors are themselves the subject of adaptive evolution, this discovery has brought into question the random nature of mutagenesis. It would be highly adaptive for organisms inhabiting variable environments to modulate mutational dynamics in ways likely to produce necessary adaptive mutations in a timely fashion while limiting the generation of other, probably deleterious, mutations."24 Evidence for the Adaptive Evolution of Mutation Rates. Minireview by David Metzgar, Christopher Wills (2000) Cell 101, p581.
The best-characterized variable genes are those used to make immunoglobulins like antibodies, which have been studied for over thirty years.25 Antibodies are proteins that are secreted into the bloodstream where they seek out and mark specific foreign substances for destruction. The ability of a limited number of these genes to produce a seemingly unlimited number of proteins has been a source of speculation for many decades. Research over the last few years has been very informative, and the cell's ability to continuously generation new alleles for this purpose is now evident. We now know that antibody specificity is due primarily to variability within the region of the protein that is involved with antigen binding. A portion of the antibody remains constant, while the antigen-binding site of the antibody is highly variable. It was discovered some time ago that the functional antibody gene is assembled during the development of the B-cell from hundreds of potential segments that lie up to a million base pairs apart. It was assumed that the production of vast numbers of antibodies was accomplished by this alone, until sequence data showed that the variable region of the gene possessed additional diversity not found in original segments. Because geneticists today do not believe the cell was designed to purposefully change genes, the source of these additional sequence alterations was originally attributed to inaccurate splicing of the various gene segments. Today it is now understood that the changes occur in a post cleavage step through the process of gene conversion.26 The variable regions of immunoglobulin genes were first demonstrated to be participating in intrachromosomal gene conversion in chickens, and these reactions are now known to be the mechanism responsible for immunity in most mammals.27 A clear connection was finally established in 2002 between these changes and the process of gene conversion when an enzyme called AID (activation-induced deaminase), which is required for antibody hypervariability, was shown to also be necessary for gene conversion in mutant cell lines.28 Two separate forms of gene conversion appear to be involved, but the exact mechanisms are not yet understood. Both types of editions utilize repetitive modifications to develop what is called immunity maturity. One form makes use of homologous pseudogenes as donor template sequences15, however, in some vertebrates including humans, the V-region is primarily edited through a system that generates single nucleotide substitutions. These findings support that even individual base pair changes in antibody genes are the result of intentional HR instead of random mutations as formerly thought.
Although the variability found in the immunoglobulin gene family is not inherited, they will likely remain the best characterized of all the variable genes, and therefore serve as a molecular model for the methods that cells use to produce new alleles. A great many heritable genes also contain regions of exceptional variability and the mechanisms responsible appear remarkable similar.29 The most polymorphic known to date also possesses an important immunological function in the recognition of self versus foreign cells, and is responsible for tissue rejection following transplant surgery. After obtaining the consensus genetic sequence of the MHC loci in 1999, well more than 100 alleles have already been found for most of the locus within this gene family. The genes of the major histocompatibility complex (MHC) are similarly edited during meiosis through the process of gene conversion using template DNA that resides elsewhere in the genome.24 By editing the genes of the MHC, new alleles are created for each offspring giving the immunity system an ability to recognize its cells apart from any other. As with all known hypervariable genes, MHC editions are not random changes. The frequency of gene conversion events varies greatly from one allele to the next, and is localized to specific areas that are rich in CpG nucleotide dimers.30
Protein toxins possessed by most animals are also hypervariable including scorpions, snakes, and cone snails.31 One particular example is the gene that makes venom for the carnivorous cone shells (Genus Conus). Conotoxins block neurotransmitters so rapidly this snail is able to use them to catch fish, and also immobilize other mollusks before they can retreat into their shell. The specificity of the toxins for ion channels makes them a valuable pharmacological tool for the treatment of neural disorders such as epilepsy. When it was recognized that conotoxins are highly diverse in their activity, it led to intense research into the genetic variability within the locus. One hundred seventy conopeptides were recently sequenced, and it was estimated that there are 100 unique peptides for each species with an estimated 50,000 present in the genus.23 Sequence analysis has revealed several nonrandom patterns that indicate the presence of a mechanism responsible for the variability. For instance, cysteine codons were found at specific positions that remain conserved within the most variable portion of the gene.22 The nucleotide substitutions that occurred were most typically nonsynonymous, and there was a bias for transversions (AT / TA) over transitional exchange. Researchers believe the striking cysteine position conversation is a molecular signature of an editing mechanism, and similar to those in the immunoglobulin loci, are required to explain the diversity in conotoxins.23 It is extremely interesting that hypervariability appears common in genes that are involved with direct interface between organisms. In addition to protein toxins, the genes used to make antitoxins are also highly variable.32 The usefulness of toxins for defense or to acquire prey is critical for survival in a predatory environment. Because these genes are variable, it would appear true that many organisms today owe their success to a skillfully designed gene editing machinery, rather than mutations or resorting of genes as previously thought. Adapting to life in a food chain appears much like ongoing warfare between two creatures that must constantly upgrade their genetic arsenal to survive. In addition to toxin and antitoxins, the genes that code for antigens in pathogenic organisms are also typically variable in order to counter antibody function. For example, the antigenic genes of the parasitic trypanosome are changed continually during infection, and are altered abruptly without increased rates of mutation elsewhere in the genome.24 Discussion It is truly more logical to propose that cells have been designed to perform some level of genetic engineering, than suggest random mutations are responsible for finely tuned adaptations. Atheists must believe that mutations are the ultimate source of new genetic information out of theoretic necessity. However, since Gregor Mendel discovered the basics of genetics, it has remained reasonably obvious that cellular mechanisms are generating the variations of related species that Darwin described. Although Mendelian genetics and selective breeding histories have answered a great portion of the mystery behind genetic diversity, the exact relationship between HR and population variety remains far from understood. Given our history of selective breeding and the apparent ability of the molecular machinery to rapidly create new alleles, it is appropriate to postulate that organisms are able to continuously produce genetic diversity. Arguably, the wolf did not already possess the variability we now find among the domestic dogs. Instead, diversity began to increase following domestication, which effectively removed the selective pressures that kept the animal true to form. It would appear that selection must persistently remove new alleles to keep a bloodline pure. That assertion is adequately demonstrated by the need to continuously remove variants from registered breeds in order to maintain desired traits. Unfortunately, the creation science community has been denying the existence of new alleles rather than looking to cellular mechanisms for the source.4 Before the rapid production of diversity can be understood from an intelligent design standpoint, we must first acknowledge that new alleles are accumulating within recognized baramin, and closely investigate these changes. Biblical references provide us adequate assurance that the alleles responsible for variety did not accumulate over millions of years by random mutations. It is also clear that there are a great many more alleles present today than could have been possessed by the population preserved from the flood. We should remember that adaptation to a particular habitat or niche involves largely uncharacterized modifications of the genome, and much of what we've learned about genetic heredity has come from theorists who do not believe the cell was designed to perform such changes with intent. The ability of the cell to produce new alleles has probably remained misunderstood for so long because the products of these reactions are being attributed to a source that is independent of cellular purpose (mutations). These assumptions overlook the fact that HR frequently demonstrates the ability to systematically produce certain outcomes. For example, vaccinations can completely eradicate disease from a population because every single individual will develop immunity if inoculated with a functional serum. There are many examples of viruses such as Polio and Small Pox that have been eliminated from the modernized world because it is unquestionable that functional antibodies will be assembled following exposure to almost any foreign substance. Since random genetic changes will simply not result in an expected sequence, the immunity system provides an excellent example of the seemingly unlimited potential of HR to generate new information. The mechanisms behind this type of gene conversion are not yet understood, but clearly illustrate the ability of the cell to specifically edit genes, and thereby rapidly multiply the number of alleles in a population. Further characterization should prove to be valuable evidence that cellular design governs the production of genetic variability, and adaptive evolution that occurs as a result.
References
|
||||||||
Receive Event
Announcements
Live Classes

At
Cedar Park Church
and Evergreen Baptist